top of page
Publications
44. Li, J.; Clark, V.; Yu, C-H.; Scida, K.; Pellitero, M. A.; Albarracín Rivera, R. L.; Zhong, W.; Demek, E.; Fountain, J.; Mahlum, J. D.; Haaland, R. E.; Carr, G. V.; Sczepanski, J. T.* and Arroyo-Currás, N.* Monitoring HIV Antiretroviral Therapy via Aptamer-Based Measurements in Preclinical Animal Models and in Human Plasma. Advanced Sensors Research 2025, 4, 2400191.

26. Kabza, A. M.; Sczepanski, J. T.* L-DNA-Based Catalytic Hairpin Assembly Circuit. Molecules 2020, 25, 947. https://doi.org/10.3390/molecules25040947

21. Banerjee, D. R.; Deckard, C. E. III; Zeng, Y.; Sczepanski, J. T.* Acetylation of the Histone H3 Tail Domain Regulates Base Excision Repair on Higher-Order Chromatin Structures. Sci. Rep. 2019, 9, 15972. https://doi.org/10.1038/s41598-019-52340-0

48. Liu, Z.; Xi, S.; McGregor, L. A.; Yamatsugu, K.; Kawashima, S.; Sczepanski, J. T.*; Kanai, M.* Constructing Nucleosome Arrays with Variable Spatial Arrangements of Histone PTMs and DNA Damage via Abiotic-Enzymatic Hybrid Catalysts and Plug-and-Play Systems. Angewandte Chemie International Edition 2025, 64, e202500162.

Before Texas A&M
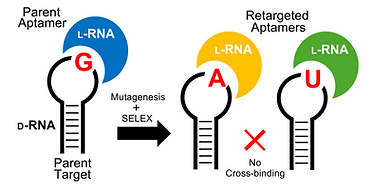
14. Sczepanski, J. T.; Joyce, G. F.* Specific Inhibition of MicroRNA Processing Using L-RNA Aptamers. J. Am. Chem. Soc. 2015, 137, 16032–16037.
13. Sczepanski, J. T.; Joyce, G. F.* A Cross-Chiral RNA Polymerase Ribozyme. Nature 2014, 515, 440–442.
12. Sczepanski, J. T.; Joyce, G. F.* Binding of a Structured -RNA Molecule by an -RNA Aptamer. J. Am Chem. Soc. 2013, 135, 13290–13293.
11. Zhou, C.; Sczepanski, J. T.; Greenberg, M. M.* Histone Modification via Rapid Cleavage of C4-Oxidized Abasic Sites in Nucleosome Core Particles. J. Am. Chem. Soc. 2013, 135, 5274–5277.
10. Sczepanski, J. T.; Zhou, C.; Greenberg, M. M.* Nucleosome Core Particle-Catalyzed Strand Scission at Abasic Sites. Biochemistry 2013, 52, 2157–2164.
9. Sczepanski, J. T.; Joyce, G. F.* Synthetic Evolving Systems that Implement a User-Specified Genetic Code of Arbitrary Design. Chem. Biol. 2012, 19, 1324–1332.
8. Zhou, C.; Sczepanski, J. T.; Greenberg, M. M.* Mechanistic Studies on Histone Catalyzed Cleavage of Apyrimidinic/Apurinic Sites in Nucleosome Core Particles. J. Am. Chem. Soc. 2012, 134, 16734–16741.
7. Sczepanski, J. T.; Hiemstra, C.; Greenberg, M. M.* Probing DNA Interstrand Cross-Link Formation by an Oxidized Abasic Site Using Nonnative Nucleotides. Bioorg. Med. Chem. 2011, 19, 5788–5793.
6. Sczepanski, J. T.; Wong, R. S.; McKnight, J. N.; Bowman, G. D.; Greenberg, M. M.* Rapid DNA-Protein Cross-Linking and Strand Scission by an Abasic Site in a Nucleosome Core Particle. Proc. Natl. Acad. Sci. USA. 2010, 107, 22475–22480.
5. Wong, R. S.; Sczepanski, J. T.; Greenberg, M. M.* Excision of a Lyase-Resistant Oxidized Abasic Lesion from DNA. Chem. Res. Toxicol. 2010, 23, 766–770.
4.Greenberg, M. M.*; Newman, C. A.; Resendiz, M.; Sczepanski, J. T. Photochemical Generation and Reactivity of 5,6-Dihydrouridin-6-yl Radical. J. Org. Chem. 2009, 74, 7007–7012.
3. Sczepanski, J. T.; Jacobs, A.; Van Houten, B.; Greenberg, M. M.* Double-Strand Break Formation During Nucleotide Excision Repair of a DNA Interstrand Cross-Link. Biochemistry 2009, 48, 7565–7567.
2. Sczepanski, J. T.; Jacobs, A.; Majumdar, A.; Greenberg, M. M.* Scope and Mechanism of Interstrand Cross-Link Formation by the C4-Oxidized Abasic Site. J. Am. Chem. Soc. 2009, 131, 11132–11139.
1. Sczepanski, J. T.; Jacobs, A.; Greenberg, M. M.* Self-Catalyzed DNA Interstrand Cross-Link Formation by an Abasic Site. J. Am. Chem. Soc. 2008, 130, 9646–9647.
bottom of page